Mfl581
formamidopyrimidine-DNA glycosylaseBBF10K_001813
source
Mesoplasma florum L1
Involved in base excision repair of DNA damaged by oxidation or by mutagenic agents. Acts as DNA glycosylase that recognizes and removes damaged bases. Has a preference for oxidized purines, such as 7,8-dihydro-8-oxoguanine (8-oxoG). Has AP (apurinic/apyrimidinic) lyase activity and introduces nicks in the DNA strand. Cleaves the DNA backbone by beta-delta elimination to generate a single-strand break at the site of the removed base with both 3'- and 5'-phosphates.
attr.
Keoni Gandall
Usage
growth
shipping strain
{shipping_strain}
growth conditions
37 C, shaking 300 rpm
antibiotic
ampicillin
expression
strain
N/A
promoter
N/A
inducer
N/A
cloning
method
GoldenGate
enzyme
BsaI
overhangs
3' - AATG … GCTT - 5'
sequencing
forward primer
M13 For
reverse primer
M13 Rev
Construct
plasmid name
pOpen-Mfl581
plasmid size
2891
insert size
828
origin
ColE1 High Copy
copy number
500-700
Safety
BSL
BSL1
other information
No Value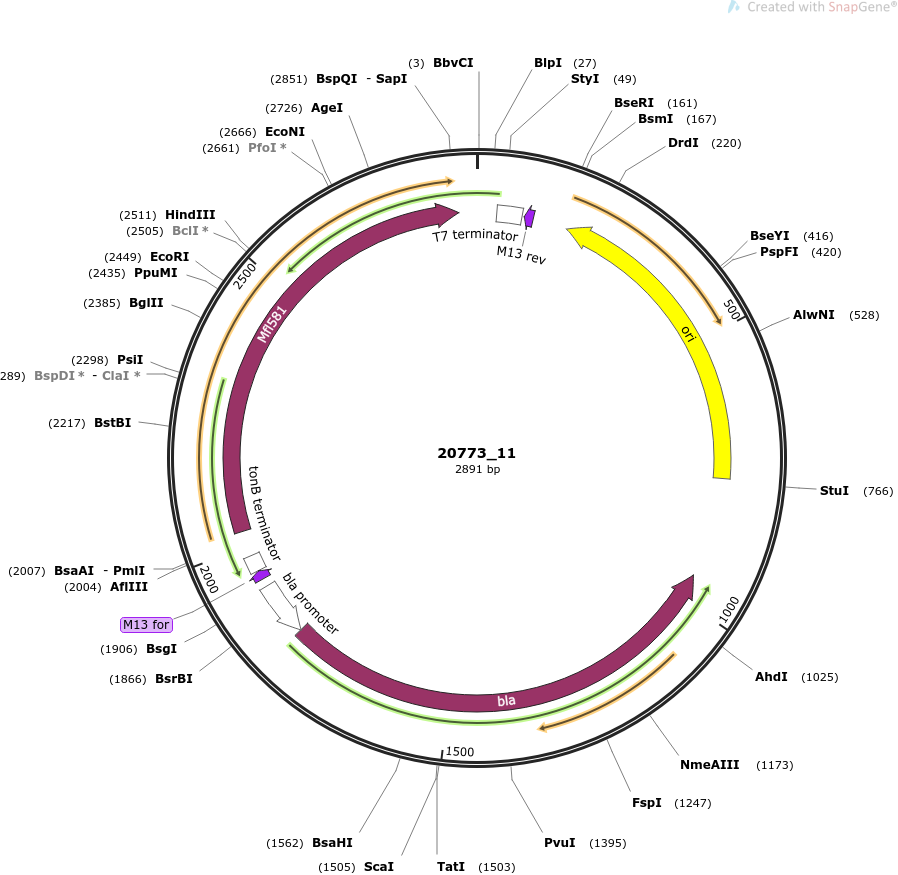
References
Available Elsewhere
FALSE